Mining millions of proteins could become faster and easier with a new technique that may also transform the enzyme-catalyst industry, according to University of California, Davis, researchers.
The research describes a strategy for quickly finding specific enzymes among the millions of proteins listed in genetic databases. The study, published Nov. 24 in the online, open-access journal Nature Communications, resulted from a collaborative research effort that included scientists from UC Davis, UCLA and the University of Washington.
The research team hopes to apply the breakthrough approach to bioengineering problems in food, health and biofuels.
"We can skip the traditional steps and jump really quickly to the enzyme of interest," said senior author Justin Siegel, a UC Davis assistant professor of chemistry and biochemistry.
The researchers filed for a patent claim on the study results — a small set of residues that clearly correlate with a targeted protein function. Residues are another term for amino acids, the building blocks of proteins. The residues identified in the study increase production of E. coli bacteria.
New technique strengthens protein patent process
"It's a completely new way of filing a patent for protein," Siegel said. Patenting residues instead of entire proteins could minimize challenges from competitors, Siegel said. "Our approach offers better protection.
"If the residues are present in any protein in this context, then we own it," Siegel said. "If you can really patent a protein well, it makes it much easier for companies to get something of value with a single patent."
Enzymes are the backbone of biology's chemical factories. These protein molecules speed up biological chemical reactions and are used to manufacture everything from medicine to biofuels. The specialized enzyme industry is worth millions of dollars, with the market estimated to hit nearly $1 billion by 2020, according to Allied Market Research.
New enzyme discovered
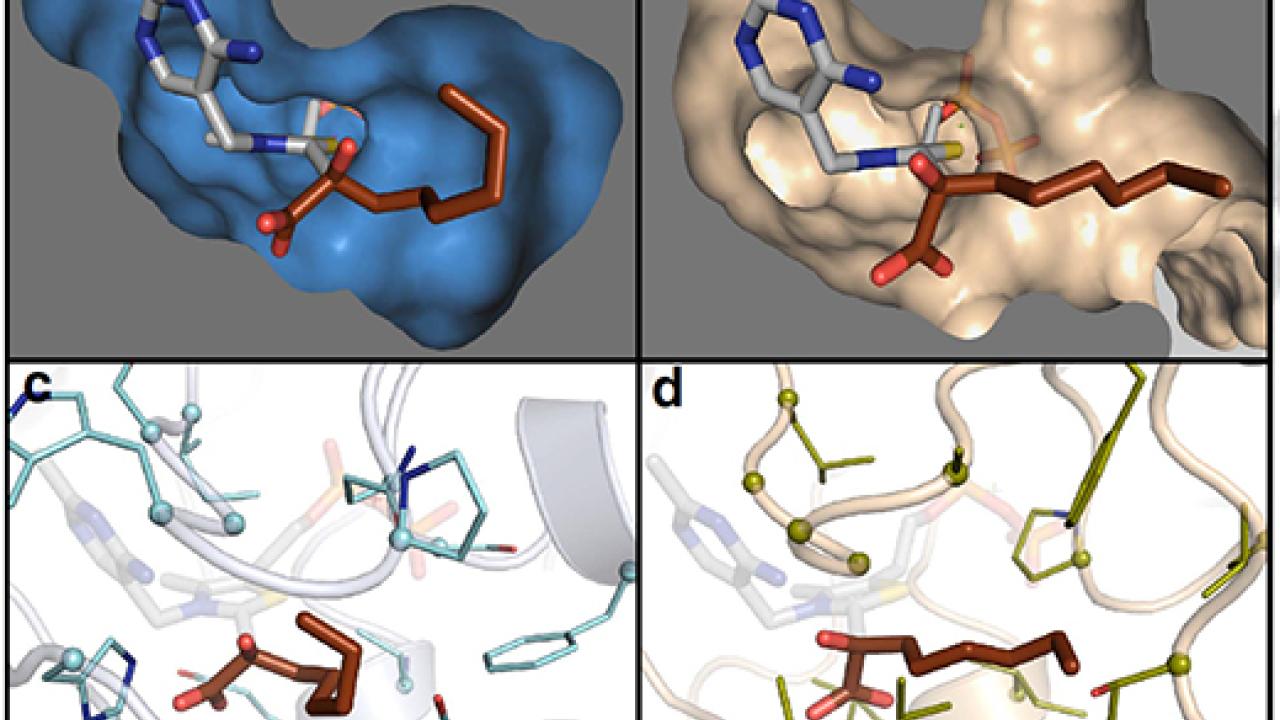
Siegel and his research colleagues discovered an enzyme that changes a key step in an E. coli metabolic pathway, which makes short-chain alcohols such as ethanol. Previous research by project collaborators at UCLA revealed that tweaking a single enzyme, called KIVD, can shift the output toward long-chain alcohols.
To find a replacement for KIVD, the UC Davis-led team used computational tools and molecular modeling to identify potential proteins listed in genome databases.
The best candidate from their search was surprisingly different from the KIVD protein, Siegel said. The two proteins shared only 15 percent of the same amino acid sequences. This is unusual because most enzyme engineering — and patents — target proteins that are 80 to 90 percent similar to the original, he said.
When lead author Wilson Mak, a UC Davis chemistry graduate student, swapped out KIVD for the new enzyme, called GEO 175, production of long-chain alcohols increased tenfold, the researchers reported. More than 95 percent of the total product consisted of long-chain alcohols such as octanol.
However, even though the protein swap pumped up long-chain alcohol production, the bacteria's overall output dropped. That's because the long-chain alcohols were toxic to the microbes, destroying their cell walls, Siegel said. Alternative production methods such as continuous extraction could solve the toxicity problem, or engineering the cells to better withstand the chemicals.
Comparing new and old techniques
The researchers also tested their new approach against a more traditional enzyme engineering technique. With this technique, the team mutated the activity of a well-known enzyme to get the desired function. This required two successive rounds of screening more than 400 mutants to find a good match.
"We had to make thousands of genes to do mutagenesis, and it took a lot more time, money and effort than genomic mining," Siegel said. "Our new methods will hopefully translate into better, cheaper, more sustainable catalysts and chemicals."
Other study authors include Stephen Tran, Ryan Marcheschi and James Liao, UCLA; Steve Bertolani, UC Davis; and James Thompson and David Baker, University of Washington.
Media Resources
Andy Fell, Research news (emphasis: biological and physical sciences, and engineering), 530-752-4533, ahfell@ucdavis.edu
Justin Siegel, UC Davis Department of Chemistry, (530) 752-9910, jbsiegel@ucdavis.edu
Becky Oskin, Mathematical and Physical Sciences, (530) 754-2222, bcoskin@ucdavis.edu